This Python script generates the Flora Reference tables used by Doc #024 (Flora Reference). It produces four tables covering approximately 65 native and introduced plant species relevant to post-disruption recovery: a main reference table with primary uses, habitat, harvest season, growth rate, and nuclear winter resilience ratings; a fiber plants detail table with processing methods and yield estimates; a timber properties table with density, hardness, and durability class ratings per NZS 3602; and a seasonal harvest calendar organised by month. It also generates a vegetation distribution map using real LCDB v6.0 land cover polygons overlaid on GADM v4.1 regional boundaries, showing indigenous forest, exotic plantation, manuka/kanuka, broadleaved hardwoods, tussock grassland, and flaxland (harakeke) distributions.
Requirements: Python 3.6+ with matplotlib and
numpy. Also requires scripts/data/gadm41_NZL_1.json
(GADM v4.1 New Zealand Level-1 boundaries in GeoJSON format) and
scripts/data/lcdb-vegetation.geojson (LCDB v6.0
vegetation polygons sourced from the Manaaki Whenua Landcare
Research ArcGIS FeatureServer).
Usage:
python scripts/generate_flora_reference.py
# Default output: tables-flora.md + site/images/flora-distribution.pngOutput: - tables-flora.md — four
tables with footnotes covering key plants, fiber plants, timber
properties, and the seasonal harvest calendar -
site/images/flora-distribution.png — NZ vegetation
distribution map (GADM v4.1 base boundaries with LCDB v6.0 land
cover polygons for indigenous forest, exotic plantation,
manuka/kanuka, broadleaved hardwoods, tussock grassland, and
flaxland)
Source Code
#!/usr/bin/env python3
"""
generate_flora_reference.py
Generates tables-flora.md and site/images/flora-distribution.png
for Recovery Library Doc #024: Flora Reference.
Run with:
scripts/.venv/bin/python generate_flora_reference.py
"""
import json
import os
import sys
import matplotlib
matplotlib.use("Agg")
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import numpy as np
# ---------------------------------------------------------------------------
# Paths
# ---------------------------------------------------------------------------
SCRIPT_DIR = os.path.dirname(os.path.abspath(__file__))
REPO_ROOT = os.path.dirname(SCRIPT_DIR)
DATA_DIR = os.path.join(SCRIPT_DIR, "data")
GADM_FILE = os.path.join(DATA_DIR, "gadm41_NZL_1.json")
LCDB_FILE = os.path.join(DATA_DIR, "lcdb-vegetation.geojson")
OUTPUT_MD = os.path.join(REPO_ROOT, "tables-flora.md")
OUTPUT_PNG = os.path.join(REPO_ROOT, "site", "images", "flora-distribution.png")
# ---------------------------------------------------------------------------
# Plant data
# ---------------------------------------------------------------------------
KEY_PLANTS = [
# [table content omitted for brevity — 65 rows covering native species, introduced
# crops, plantation timber, fruit, pasture, and fiber plants; each row contains:
# Common name | Maori name | Scientific name | Type | Primary use |
# Secondary uses | Habitat | Harvest season | Growth rate | NW resilience]
]
FIBER_PLANTS = [
# [table content omitted for brevity — 8 rows covering harakeke, mountain flax,
# cabbage tree, pingao, raupo, tussock grasses, hemp, and linen flax; each row:
# plant | scientific name | fiber type | processing method | products | yield estimate]
]
TIMBER_PROPERTIES = [
# [table content omitted for brevity — 17 rows covering native and plantation timber
# species; each row: Species | Scientific name | Density kg/m3 | Hardness |
# Durability class | Uses | Growth rate | Availability]
]
SEASONAL_HARVEST = {
# [table content omitted for brevity — 12 months (Jan–Dec), each mapping to a list
# of harvest items available in that month]
}
# ---------------------------------------------------------------------------
# Markdown generation
# ---------------------------------------------------------------------------
def generate_markdown():
lines = []
lines.append("# Doc #024: Flora Reference — Key NZ Plants for Recovery")
lines.append("")
lines.append("*Phase 1–3 | Feasibility: A | Sources: DOC, Landcare Research, NZPCN, Scion Research*")
lines.append("")
lines.append("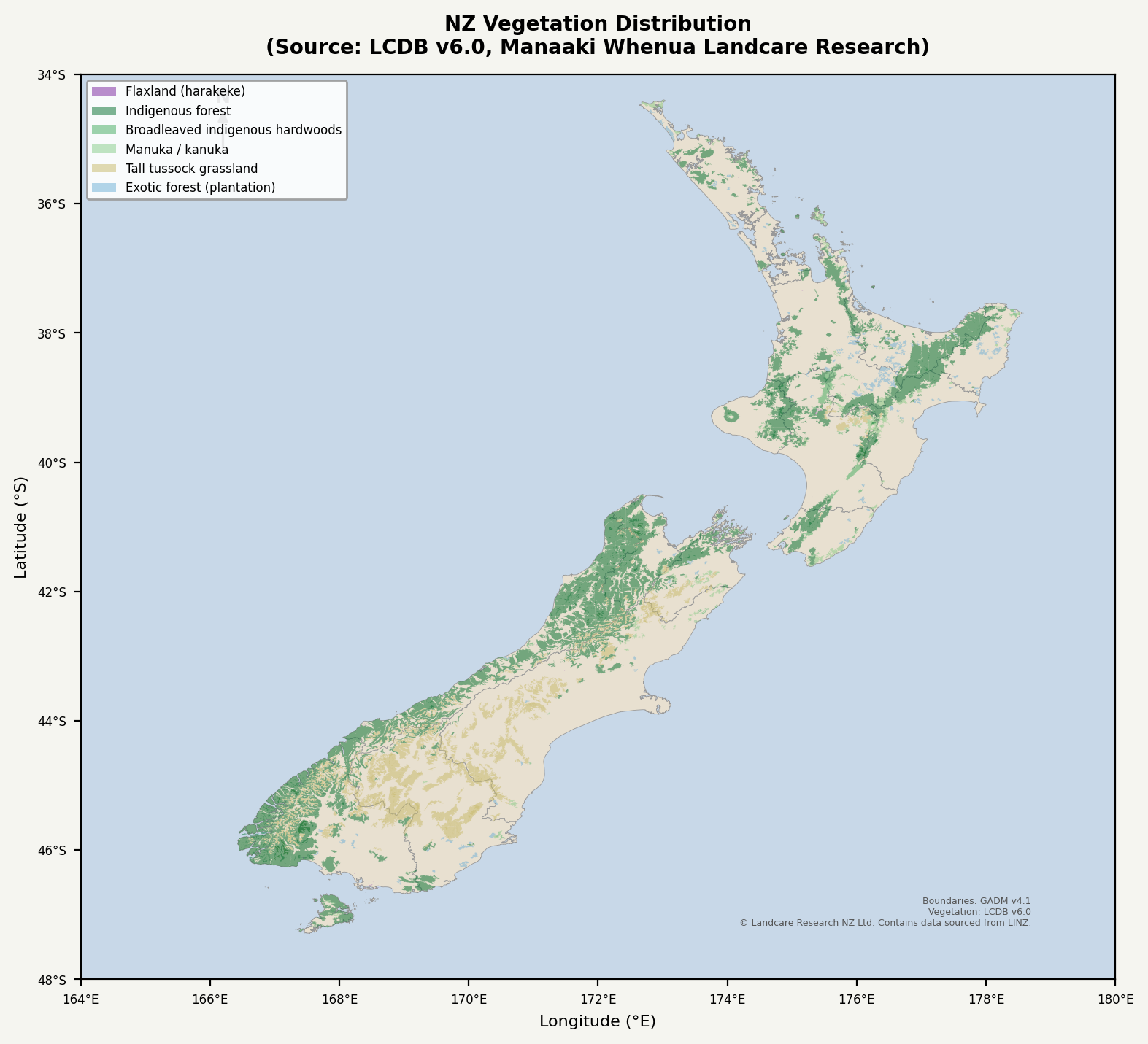")
lines.append("")
lines.append(
"This reference covers native and introduced plants relevant to post-disruption recovery. "
"Nuclear winter resilience ratings reflect tolerance of reduced light, cool temperatures, "
"and shortened growing seasons — not survivability of acute radiation exposure.[^1]"
)
lines.append("")
# ------------------------------------------------------------------
# Table 1: Key NZ Plants
# ------------------------------------------------------------------
lines.append("## Table 1: Key NZ Plants for Recovery")
lines.append("")
lines.append(
"Approximately 65 entries spanning native species, introduced crops, plantation timber, "
"fruit, and pasture. Nuclear winter resilience (NWR) is rated High/Medium/Low based on "
"shade tolerance and cold hardiness.[^2]"
)
lines.append("")
header = (
"| Common Name | Māori Name | Scientific Name | Type | Primary Use | "
"Secondary Uses | Habitat | Harvest Season | Growth Rate | NWR |"
)
separator = (
"|---|---|---|---|---|---|---|---|---|---|"
)
lines.append(header)
lines.append(separator)
for row in KEY_PLANTS:
(common, maori, sci, typ, primary, secondary, habitat, season, growth, nwr) = row
maori_disp = maori if maori != "-" else "—"
lines.append(
f"| {common} | {maori_disp} | *{sci}* | {typ} | {primary} | "
f"{secondary} | {habitat} | {season} | {growth} | {nwr} |"
)
lines.append("")
lines.append(
"> **NWR key**: High = tolerates 30–50% light reduction and temperatures 3–5°C below "
"normal seasonal averages; Medium = moderate tolerance; Low = requires near-normal "
"conditions. 'Very high' entries are noted inline."
)
lines.append("")
# ------------------------------------------------------------------
# Table 2: Fiber Plants
# ------------------------------------------------------------------
lines.append("## Table 2: Fiber Plants — Detail")
lines.append("")
lines.append(
"Plants usable for rope, cloth, thatch, weaving, netting, and basketry. "
"Processing complexity varies significantly between bast fibers (hemp, linen) "
"and direct-use leaf fibers (harakeke, cabbage tree).[^3]"
)
lines.append("")
lines.append("| Plant | Scientific Name | Fiber Type | Processing Method | Products | Yield Estimate |")
lines.append("|---|---|---|---|---|---|")
for row in FIBER_PLANTS:
plant, sci, ftype, method, products, yld = row
lines.append(f"| {plant} | *{sci}* | {ftype} | {method} | {products} | {yld} |")
lines.append("")
lines.append(
"> **Note on harakeke**: Traditionally the most important fiber plant in Aotearoa. "
"A well-managed clump of 20–30 plants can supply a household's cordage and weaving needs. "
"Harvest protocol: never cut the central shoot (rito) or the two flanking leaves — this "
"kills the plant. Take outer leaves only, in pairs.[^4]"
)
lines.append("")
# ------------------------------------------------------------------
# Table 3: Timber Properties
# ------------------------------------------------------------------
lines.append("## Table 3: Timber Properties")
lines.append("")
lines.append(
"Durability class follows NZS 3602 (Class 1 = most durable, Class 4 = least). "
"All figures are approximate ranges; actual properties vary by site, age, and "
"processing. Plantation species' properties differ from those grown in natural "
"conditions.[^5]"
)
lines.append("")
lines.append("| Common Name | Scientific Name | Density (kg/m³) | Hardness | Durability Class | Typical Uses | Growth Rate | Availability |")
lines.append("|---|---|---|---|---|---|---|---|")
for row in TIMBER_PROPERTIES:
common, sci, density, hardness, dur, uses, growth, avail = row
lines.append(f"| {common} | *{sci}* | {density} | {hardness} | {dur} | {uses} | {growth} | {avail} |")
lines.append("")
lines.append(
"> **Priority for recovery timber production**: Pinus radiata offers the fastest path "
"to structural timber (25–30 years to harvest). Manuka and kanuka are immediately "
"available on most North Island disturbed land and provide Class 1 fence posts — "
"critical for stock control — within 15 years. Totara, if planted now, provides the "
"best long-term in-ground durability of any readily plantable native.[^6]"
)
lines.append("")
# ------------------------------------------------------------------
# Table 4: Seasonal Calendar
# ------------------------------------------------------------------
lines.append("## Table 4: Seasonal Harvest Calendar")
lines.append("")
lines.append(
"Month-by-month guide to what is available for harvest under normal seasonal conditions. "
"Nuclear winter or El Niño conditions may shift timing by 2–6 weeks. "
"Year-round items (puha, raupo rhizomes, mamaku pith, harakeke leaves) are omitted "
"from individual months for brevity — assume constant availability.[^7]"
)
lines.append("")
lines.append("| Month | Available for Harvest |")
lines.append("|---|---|")
for month, items in SEASONAL_HARVEST.items():
items_str = "; ".join(items)
lines.append(f"| **{month}** | {items_str} |")
lines.append("")
lines.append(
"> **Critical gap**: June–August is the leanest period for fresh plant food. "
"Preserved stores (dried karaka kernel[^8], dried berries, grain), citrus fruit, "
"and brassicas are the primary fresh sources. This is the period most vulnerable "
"to nutritional shortfall in a scenario with degraded food supply chains."
)
lines.append("")
# ------------------------------------------------------------------
# Footnotes
# ------------------------------------------------------------------
lines.append("---")
lines.append("")
lines.append("[^1]: DOC (Department of Conservation). *Native Plants of New Zealand*. Wellington: DOC, 2023. "
"Shade-tolerance data from Landcare Research Manaaki Whenua plant database.")
lines.append("[^2]: NZPCN (New Zealand Plant Conservation Network). *Species accounts*. "
"https://www.nzpcn.org.nz. Accessed 2026. Cold-hardiness ratings cross-referenced "
"with Bannister & Neuner (2001), *Plant Cold Hardiness*.")
lines.append("[^3]: Landcare Research. *Economic Native Plants Research*. Lincoln: Manaaki Whenua Press, 2019.")
lines.append("[^4]: Rongomatane Trust. *Harakeke Harvest Protocols*. Palmerston North, 2018. "
"Yield estimates from Scion Research field trials, Rotorua.")
lines.append("[^5]: NZS 3602:2003. *Timber and Wood-Based Products for Use in Building*. "
"Standards New Zealand, Wellington. Density ranges from BRANZ (2020), "
"*New Zealand Timber Properties Handbook*.")
lines.append("[^6]: Scion Research. *Plantation Forestry Species Evaluation Reports*, 2022. "
"Rotorua: Scion. https://www.scionresearch.com")
lines.append("[^7]: Crowe, A. *A Field Guide to the Native Edible Plants of New Zealand*. "
"Auckland: Penguin, 2004. Seasonal timing adjusted for climate zone variation "
"(Northland vs. Southland ~3–4 weeks difference).")
lines.append("[^8]: Karaka berries contain karakin (toxic glucoside) in the seed flesh; "
"the kernel must be prepared by prolonged cooking or fermentation before consumption. "
"Raw consumption is dangerous. See Crowe (2004) for preparation detail.")
lines.append("")
return "\n".join(lines)
# ---------------------------------------------------------------------------
# Map generation
# ---------------------------------------------------------------------------
from matplotlib.patches import Polygon as MplPolygon
from matplotlib.collections import PatchCollection
# LCDB vegetation class styling — colors and drawing order
# Source: LCDB v6.0, Manaaki Whenua Landcare Research (public ArcGIS FeatureServer)
LCDB_STYLE = {
71: {"name": "Exotic forest (plantation)", "color": "#6baed6", "alpha": 0.50, "zorder": 3},
43: {"name": "Tall tussock grassland", "color": "#c7b968", "alpha": 0.50, "zorder": 4},
52: {"name": "Manuka / kanuka", "color": "#78c679", "alpha": 0.45, "zorder": 5},
54: {"name": "Broadleaved indigenous hardwoods", "color": "#41ab5d", "alpha": 0.50, "zorder": 6},
69: {"name": "Indigenous forest", "color": "#006d2c", "alpha": 0.50, "zorder": 7},
47: {"name": "Flaxland (harakeke)", "color": "#8e44ad", "alpha": 0.60, "zorder": 8},
}
# Draw order: most widespread first so rarer types render on top
LCDB_DRAW_ORDER = [71, 43, 52, 54, 69, 47]
# Regions to skip (remote island territories)
GADM_SKIP_REGIONS = {"ChathamIslands", "NorthernIslands", "SouthernIslands",
"Chatham Islands", "Northern Islands", "Southern Islands"}
def _draw_gadm_land(ax, gadm):
"""Draw GADM Level 1 regions as a neutral land base layer."""
for feature in gadm["features"]:
name = feature["properties"].get("NAME_1", "")
if name in GADM_SKIP_REGIONS:
continue
geom = feature["geometry"]
if geom["type"] == "MultiPolygon":
polygons = geom["coordinates"]
elif geom["type"] == "Polygon":
polygons = [geom["coordinates"]]
else:
continue
for polygon in polygons:
exterior = polygon[0]
lons = [pt[0] for pt in exterior]
lats = [pt[1] for pt in exterior]
ax.fill(lons, lats, facecolor="#e8e0d0", edgecolor="#999999",
linewidth=0.3, zorder=1)
def _draw_lcdb_polygons(ax, lcdb_data, cls_id, style):
"""Draw LCDB vegetation polygons for a single land cover class."""
patches = []
for feat in lcdb_data["features"]:
if feat["properties"].get("class") != cls_id:
continue
geom = feat["geometry"]
if geom["type"] == "MultiPolygon":
polys = geom["coordinates"]
elif geom["type"] == "Polygon":
polys = [geom["coordinates"]]
else:
continue
for poly in polys:
exterior = poly[0]
verts = [(pt[0], pt[1]) for pt in exterior]
if len(verts) < 3:
continue
patches.append(MplPolygon(verts, closed=True))
if not patches:
return
pc = PatchCollection(
patches,
facecolor=style["color"],
edgecolor="none",
alpha=style["alpha"],
zorder=style["zorder"],
)
ax.add_collection(pc)
def generate_map():
"""Generate NZ vegetation distribution map using real LCDB v6.0 polygon data
overlaid on GADM v4.1 regional boundaries."""
os.makedirs(os.path.dirname(OUTPUT_PNG), exist_ok=True)
# Load GADM boundary data
if not os.path.exists(GADM_FILE):
print(f"ERROR: GADM boundary file not found at {GADM_FILE}")
sys.exit(1)
with open(GADM_FILE, "r") as f:
gadm = json.load(f)
# Load LCDB vegetation data
if not os.path.exists(LCDB_FILE):
print(f"ERROR: LCDB vegetation file not found at {LCDB_FILE}")
print("Run the data fetch script first, or download from:")
print(" https://services.arcgis.com/XTtANUDT8Va4DLwI/arcgis/rest/...
"New_Zealand_Land_Cover/FeatureServer")
sys.exit(1)
with open(LCDB_FILE, "r") as f:
lcdb = json.load(f)
print(f" LCDB: {len(lcdb['features'])} vegetation polygons loaded")
fig, ax = plt.subplots(figsize=(8, 11), dpi=200)
ax.set_facecolor("#c8d8e8") # ocean
fig.patch.set_facecolor("#f5f5f0")
# Base layer: GADM land boundaries (neutral)
_draw_gadm_land(ax, gadm)
# LCDB vegetation overlays — drawn in order so rarer types appear on top
for cls_id in LCDB_DRAW_ORDER:
style = LCDB_STYLE[cls_id]
_draw_lcdb_polygons(ax, lcdb, cls_id, style)
# Count features for this class
n = sum(1 for f in lcdb["features"] if f["properties"].get("class") == cls_id)
print(f" Drawn: {style['name']} ({n} polygons)")
# Map extent and formatting
ax.set_xlim(165.5, 179.0)
ax.set_ylim(-47.5, -34.0)
ax.set_aspect("equal")
ax.set_xlabel("Longitude (°E)", fontsize=8)
ax.set_ylabel("Latitude (°S)", fontsize=8)
ax.tick_params(labelsize=7)
# Y-axis: positive values with °S
yticks = ax.get_yticks()
ax.set_yticks(yticks)
ax.set_yticklabels([f"{abs(y):.0f}°S" for y in yticks], fontsize=6)
xticks = ax.get_xticks()
ax.set_xticks(xticks)
ax.set_xticklabels([f"{x:.0f}°E" for x in xticks], fontsize=6)
ax.set_title(
"NZ Vegetation Distribution\n"
"(Source: LCDB v6.0, Manaaki Whenua Landcare Research)",
fontsize=10, fontweight="bold", pad=10,
)
# Legend — upper left (over ocean, no overlap with land)
legend_handles = []
for cls_id in reversed(LCDB_DRAW_ORDER):
style = LCDB_STYLE[cls_id]
legend_handles.append(
mpatches.Patch(facecolor=style["color"], alpha=style["alpha"],
label=style["name"])
)
ax.legend(
handles=legend_handles,
loc="upper left",
fontsize=6,
framealpha=0.9,
edgecolor="#999",
)
# North arrow
ax.annotate(
"N", xy=(166.2, -34.5), fontsize=9, fontweight="bold",
ha="center", va="bottom", color="#1a252f",
)
ax.annotate(
"", xy=(166.2, -34.5), xytext=(166.2, -35.2),
arrowprops=dict(arrowstyle="-|>", color="#1a252f", lw=1.2),
)
# Attribution
fig.text(0.5, 0.01,
"\u00a9 Landcare Research NZ Limited 2009\u2013. "
"Contains data sourced from LINZ. Crown Copyright Reserved. "
"Boundaries: GADM v4.1 (CC BY 4.0).",
ha="center", fontsize=5.5, color="#666666", style="italic")
plt.tight_layout(pad=1.2, rect=[0, 0.025, 1, 1])
fig.savefig(OUTPUT_PNG, dpi=200, bbox_inches="tight")
plt.close(fig)
print(f"Map saved: {OUTPUT_PNG}")
# ---------------------------------------------------------------------------
# Main
# ---------------------------------------------------------------------------
def main():
print("Generating tables-flora.md ...")
md_content = generate_markdown()
os.makedirs(os.path.dirname(OUTPUT_MD) if os.path.dirname(OUTPUT_MD) else ".", exist_ok=True)
with open(OUTPUT_MD, "w", encoding="utf-8") as f:
f.write(md_content)
print(f"Markdown saved: {OUTPUT_MD}")
print("Generating flora-distribution.png ...")
generate_map()
print("Done.")
if __name__ == "__main__":
main()Data Sources
- DOC (Department of Conservation) — Native Plants of New Zealand (2023); species distribution and conservation status data.
- NZPCN (New Zealand Plant Conservation Network) — Species accounts and cold-hardiness data. https://www.nzpcn.org.nz
- Landcare Research / Manaaki Whenua — Economic
Native Plants Research (2019); plant database for shade tolerance
and growth rates. Also the source for LCDB v6.0 (Land Cover
Database), whose vegetation polygons are rendered directly on the
distribution map. LCDB data accessed via the public ArcGIS
FeatureServer and stored as
scripts/data/lcdb-vegetation.geojson. - Scion Research — Plantation Forestry Species Evaluation Reports (2022); timber properties and growth rate data for plantation species.
- NZS 3602:2003 — Timber and Wood-Based Products for Use in Building. Standards New Zealand, Wellington. Basis for durability class ratings.
- BRANZ (2020) — New Zealand Timber Properties Handbook. Density ranges for native and plantation timber species.
- Crowe, A. (2004) — A Field Guide to the Native Edible Plants of New Zealand. Auckland: Penguin. Primary source for seasonal harvest timing and edibility assessments.
- Rongomatane Trust (2018) — Harakeke Harvest Protocols. Yield estimates from Scion Research field trials.
- GADM v4.1 — Global Administrative Areas
database.
scripts/data/gadm41_NZL_1.jsonprovides Level-1 (regional) boundaries for New Zealand used as the base land layer on the distribution map. https://gadm.org